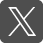DNA barcode for fungi

Dr Gareth Griffith and Dr Joan Edwards from the Institute of Biological, Environmental and Rural Sciences.
28 March 2012
Three scientists at the Institute of Biological, Environmental and Rural Sciences (IBERS) at Aberystwyth University are part of an international consortium which has agreed upon a standard ‘fungal DNA barcode'.
The conclusions of the study, which provides the foundation for the use of standardised DNA technologies for identifying fungi, are featured in the current issue of the journal PNAS (Proceedings of the National Academy of Science USA) which is published this week.
Dr Gareth Wyn Griffith, Dr Joan Edwards, and Brian Douglas from IBERS are members of a 140 strong team of scientists from more than 20 countries who collaborated on the work.
DNA barcoding is the use of a standardised region of DNA for identifying species. However, an appropriate barcode region needs to be defined for each group of living organisms. Therefore, zoologists have opted for a barcode based on mitochondrial DNA while botanists use chloroplast DNA.
Contrary to popular belief, the fungi form a distinct kingdom quite unrelated to plants, so they do not contain chloroplasts. Furthermore, some fungi are anaerobic, so mitochondrial DNA barcoding regions favoured by zoologists are also unsuitable. Having explored several options, the ITS (internal transcribed spacer) region which contains genes involved in protein synthesis was found to be the most suitable for the fungi.
The DNA barcoding technique will work on tiny amounts of material so can be used to detect fungi from air samples or from plant or animal tissues colonised by fungi. In cases of mushroom poisoning it could even be used to identify the fungus from stomach contents.
Although some fungi can produce quite large structures, for example mushrooms, most fungi are not readily visible as they grow in soil or wood. Their fine hyphal filaments are impossible to identify even with a microscope, so DNA barcoding permits huge advances in disease diagnosis and ecological studies.
The term DNA barcoding was first used in 2003 and by now the ‘barcode library’ for animals covers more than 70,000 species.
The idea for the FBOL consortium (Fungal Barcode of Life) was hatched at the International Mycological Congress in Edinburgh in 2010. It follows on from an earlier international consortium project, AFTOL (All Fungus Tree of Life), which aimed to identify natural groupings and true relatedness of all the main fungal groupings.
Dr Griffith, who was also involved in AFTOL, said, “It is wonderful to be a part of these truly global initiatives where international collaboration has facilitated very rapid progress and agreement.”
The consortium was led by Dr Conrad Schoch and Dr Keith Seifert. Dr Schoch is a taxonomist based at the National Center for Biotechnology Information (NCBI) in Bethesda Maryland which operates the Genbank DNA database, with a major part of his work being to ensure that sequences relating to fungi are properly classified and curated.
Dr Seifert is based at Agriculture and Agri-Food Canada in Ottawa and was trained by the eminent Aberystwyth alumnus Stan Hughes from Llanelli who emigrated to Canada in the early 1950’s (but is still actively naming new fungi at the age of 94).
Ironically, mycologists were the first biologists of all to undertake barcoding, way back in the early 1990’s when some of the key technologies of modern genetics such as the polymerase chain reaction (PCR) had just been invented.
Despite the fact that the Californian mycologist Tom Bruns suggested the ITS region as a standardised region for fungi in 1997, it has taken a remarkably long time for its DNA barcode status to become ‘official’ with the publication of this paper.
The Aberystwyth University team contributed to the paper through their expertise with two particular groups of fungi, the anaerobic rumen fungi and the dark septate endophytes.
The anaerobic fungi are the most primitive of all fungi and are important in breakdown of plant material in the digestive systems of sheep, cattle and other herbivores.
Dr Joan Edwards, who works on a BBSRC (Biotechnology and Biological Sciences Research Council) -funded project to study ruminal plant-microbe interactions, said “There were some discussion about using the mitochondrial DNA barcode which would not have been suitable for our anaerobic fungi, so I’m pleased that the ITS region won the day.”
Brian Douglas, a PhD student funded by the Aberystwyth Postgraduate Studentship scheme, is studying the role of dark septate endophytes in the roots of grassland plants, where they may play a role in host nutrition. “ITS sequence databases are already an amazing resource for understanding the distribution and ecology of many groups of fungi, especially those hidden away in plant tissues and soils. With the ITS region as the official fungal barcode it is now much easier for future work to build on these past few decades of molecular mycology” said Mr Douglas.