DNA metabarcoding for herbivore diet analysis
[February 2026]
This article explores a study led by Dr Hannah Vallin that was published in Ecology and Evolution in January 2026. The research was carried out in collaboration with Prof. Mariecia Fraser (IBERS), alongside colleagues from the University of Sheffield (Dr Helen Hipperson) and the James Hutton Institute (Prof. Robin Pakeman).
The study investigates the use of DNA metabarcoding, a technique that identifies which plants animals have eaten by analysing tiny fragments of plant DNA in their faeces. It works by using DNA barcodes, which are short, distinctive sections of genetic code that act like molecular “ID tags”for different plant species. By comparing these barcodes against reference databases, researchers can determine which plants were present in an animal’s diet, even when the material is too digested to be recognised visually.
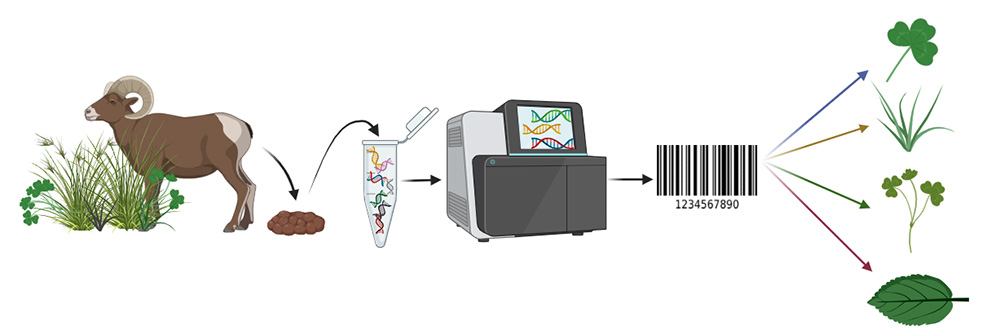
Why is this important?
Understanding what herbivores eat is important for several reasons. Diet composition affects animal health, growth and productivity, and helps land managers ensure that grazing systems remain sustainable. It also influences how herbivores shape the landscape and therefore affects biodiversity, habitat structure, and long-term ecosystem resilience. By accurately determining which plants are being consumed, researchers and land managers can:
- better understand grazing impacts,
- plan conservation strategies, and
- support evidence-based decisions in upland and lowland farming systems.
The Study
The researchers tested how accurately two commonly used plant DNA barcodes (ITS2 and trnL) reflected the actual diets of sheep fed controlled forage mixtures. They found that while the technique can reliably identify many plant species including both major and minor components of the diet, it is less accurate at estimating the proportion of each plant in the diet—especially when the forage is harder to digest. Some species, such as Medicago sativa, were detected even at very low levels (1% of the diet) but tended to be over-represented in the DNA results due to several biological biases.
These findings highlight both the strengths and limitations of DNA metabarcoding for grazing ecology. The authors emphasise the value of using multiple DNA markers and calibration controls to improve accuracy, while noting that the method already offers powerful insights into herbivore feeding patterns and plant–animal interactions.
This work directly supports IBERS’ commitment to sustainable agriculture, food security, and resilient ecosystems. By improving our ability to monitor what grazing animals eat, and how their diets change with forage availability, climate, and land‑use practices, this research strengthens our understanding of how to manage grasslands in ways that balance productivity with biodiversity and environmental stewardship. It contributes to the wider strategic aim of developing evidence-based tools that help farmers, policymakers and conservationists make informed decisions that support both profitable farming and healthy landscapes.
The paper, Evaluating the Quantitative Accuracy and Application of DNA Metabarcoding for Dietary Reconstruction in Ruminants, is available online (https://doi.org/10.1002/ece3.72878).
